
-
Careers • News • Contact us •
- Login
- Français
Creating new
knowledge
Exploring new avenues to develop tomorrow’s medical knowledge through an approach that integrates basic and clinical research
Our research units are led by principal investigators who collaborate in a spirit of collegiality and with the vision of bridging the gap between research and patients. They train the next generation of scientists and are independent and creative minds who work tirelessly to improve health.
Current research
"We work to understand how noncoding RNAs and regulatory elements in our genome impact diseases with the goal to develop novel therapies and better treatments."
Martin Sauvageau, research director
RNA and Noncoding Mechanisms of Disease
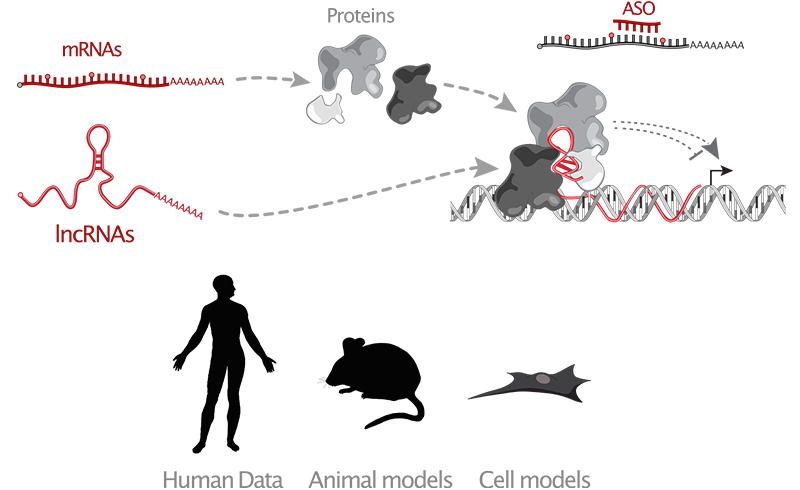
Efforts to understand the genetic and molecular interactions contributing to diseases have focused mainly on the 2% of our genome that encodes proteins, leaving the rest vastly under-explored. Yet, most genetic variations associated with human disease are found in the noncoding portions of our genome. Interestingly, a large fraction of these genomic regions is transcribed, generating tens of thousands of long RNAs devoid of protein coding potential. These long noncoding RNAs (lncRNAs) are expressed with high tissue specificity, and several were found to regulate key cellular processes through largely unknown mechanisms. Evidence also points to lncRNA loci as risk factors frequently deregulated or mutated in a wide variety of human diseases. However, the extent to which lncRNAs contribute to development in vivo and how they affect transcriptional programs and signaling pathways remain poorly characterized. Thus, one of the main challenges to understand the noncoding genome’s influence on the fundamental mechanisms of life is not only to determine which lncRNAs are functional, but also decipher how they perform their tasks.
Our laboratory combines functional genomics, proteomics and CRISPR-based genome editing techniques to perturb lncRNA functions and characterize their role at a cellular and physiological level. We also aim to understand the molecular grammar that underlies lncRNA function and uncover novel noncoding RNA-based mechanisms to develop novel molecular tools and therapies. For this, we use a combination of biochemistry, high-throughput, and computational approaches to identify RNA-interacting macromolecules and RNA domains that mediate their function.
Our goal is to better understand the impact that lncRNAs and noncoding regions have on development and reveal novel RNA-based mechanisms that could be leveraged towards their use in the clinic.
Our laboratory combines functional genomics, proteomics and CRISPR-based genome editing techniques to perturb lncRNA functions and characterize their role at a cellular and physiological level. We also aim to understand the molecular grammar that underlies lncRNA function and uncover novel noncoding RNA-based mechanisms to develop novel molecular tools and therapies. For this, we use a combination of biochemistry, high-throughput, and computational approaches to identify RNA-interacting macromolecules and RNA domains that mediate their function.
Our goal is to better understand the impact that lncRNAs and noncoding regions have on development and reveal novel RNA-based mechanisms that could be leveraged towards their use in the clinic.
| Team |
|
| Affiliations |
|
| Degrees and experience |
|
| Publications |
|
Dissecting the functions and molecular grammar of long noncoding RNAs in development and disease.
Functional Profiling
We perform functional assays to determine the role of lncRNAs in human and animal cells using genome editing, CRISPR systems and antisense oligos technologies.
Biochemistry & Genomics
We dissect the composition of RNA-based machines and characterize their molecular activities using state-of-the-art biochemical, molecular as well as short- and long-read sequencing-based approaches. We also work to develop novel RNA-based tools for targeted genome regulation.
Disease Modeling & RNA therapy
We characterize the in vivo roles of lncRNAs using animal models and xenografts combined with a suite of CRISPR-based genome engineering approaches. We also actively develop novel mRNA-based and RNA-targeting therapeutics.
Important Discoveries
Tools
| Press review |
|
| Grants |
|
| Recognitions and honors |
|
Join the
group Research
Available positions
Ongoing clinical studies
Research Partners










Join the
research group
514 987-5599
martin.sauvageau@ircm.qc.ca

© Montreal Clinical Research Institute, Année.All rights reserves. | Privacy policy | Terms of use | Web site by Agence Riposte


